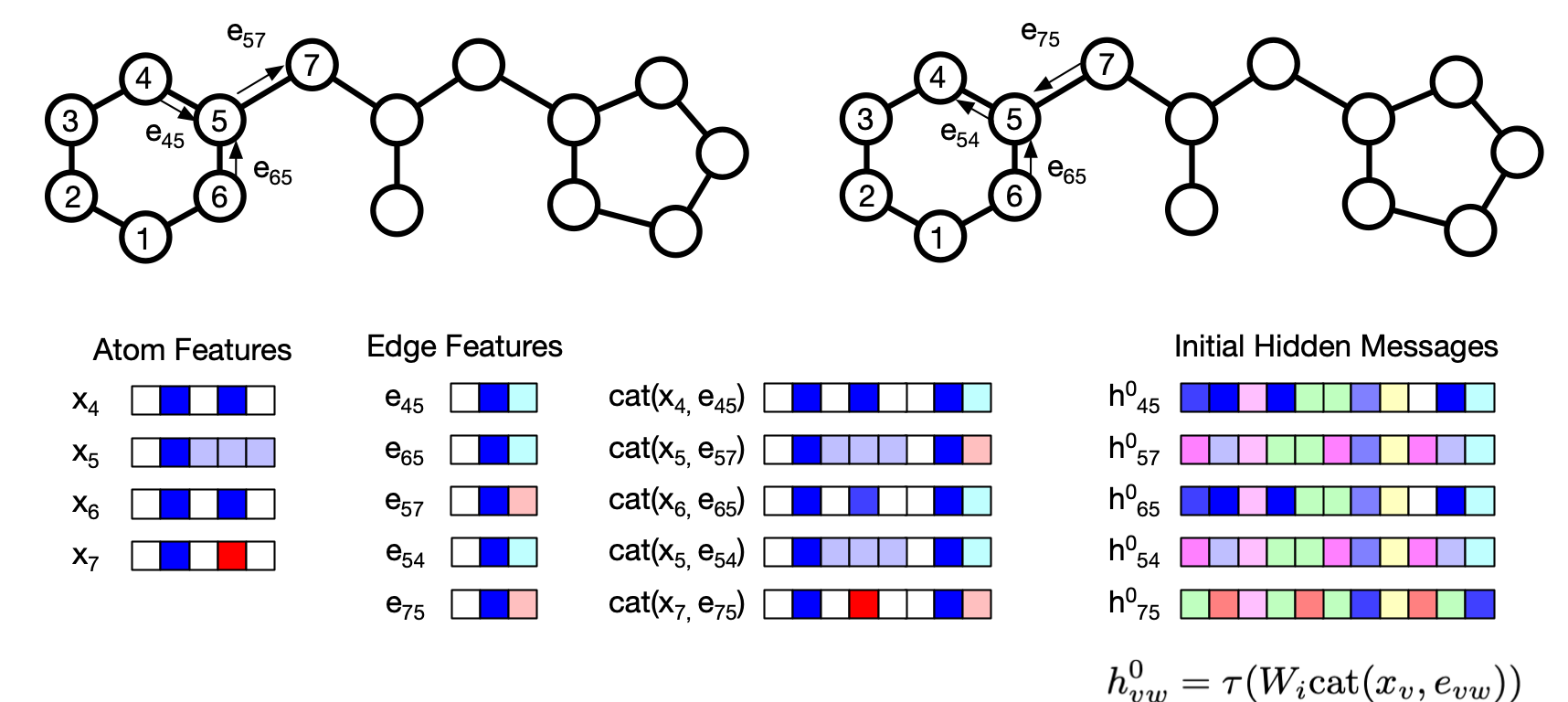
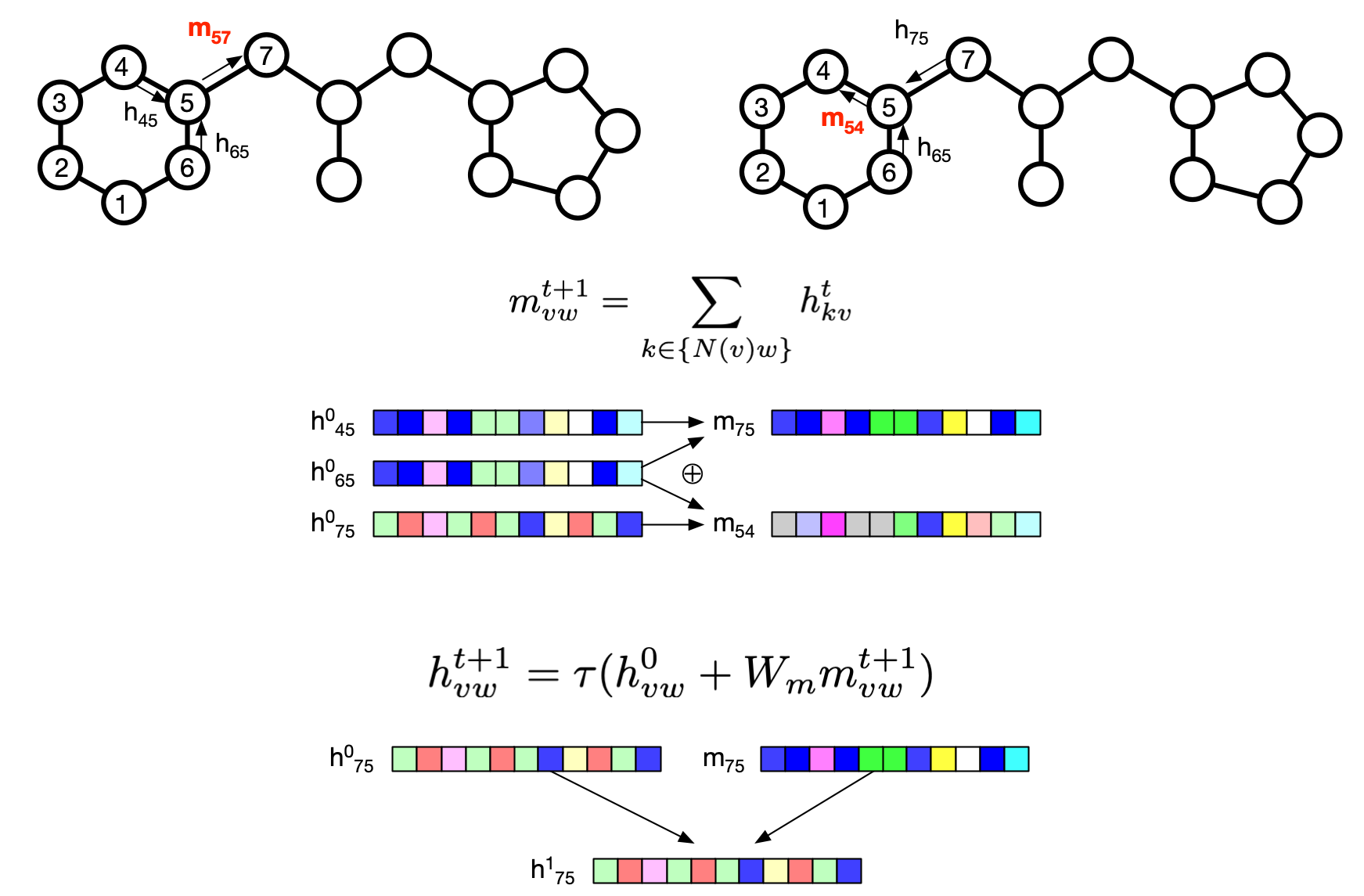
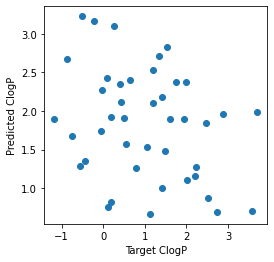
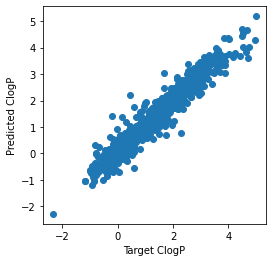
# Atom feature sizes
MAX_ATOMIC_NUM = 100
ATOM_FEATURES = {
'atomic_num': list(range(MAX_ATOMIC_NUM)),
'degree': [0, 1, 2, 3, 4, 5],
'formal_charge': [-1, -2, 1, 2, 0],
'chiral_tag': [0, 1, 2, 3],
'num_Hs': [0, 1, 2, 3, 4],
'hybridization': [
Chem.rdchem.HybridizationType.SP,
Chem.rdchem.HybridizationType.SP2,
Chem.rdchem.HybridizationType.SP3,
Chem.rdchem.HybridizationType.SP3D,
Chem.rdchem.HybridizationType.SP3D2
],
}
# Distance feature sizes
PATH_DISTANCE_BINS = list(range(10))
THREE_D_DISTANCE_MAX = 20
THREE_D_DISTANCE_STEP = 1
THREE_D_DISTANCE_BINS = list(range(0, THREE_D_DISTANCE_MAX + 1, THREE_D_DISTANCE_STEP))
# len(choices) + 1 to include room for uncommon values; + 2 at end for IsAromatic and mass
ATOM_FDIM = sum(len(choices) + 1 for choices in ATOM_FEATURES.values()) + 2
EXTRA_ATOM_FDIM = 0
BOND_FDIM = 14
def get_atom_fdim() -> int:
"""Gets the dimensionality of the atom feature vector."""
return ATOM_FDIM + EXTRA_ATOM_FDIM
def get_bond_fdim() -> int:
"""Gets the dimensionality of the bond feature vector.
"""
return BOND_FDIM + get_atom_fdim()
def onek_encoding_unk(value: int, choices: List[int]) -> List[int]:
encoding = [0] * (len(choices) + 1)
index = choices.index(value) if value in choices else -1
encoding[index] = 1
return encoding
def atom_features(atom: Chem.rdchem.Atom, functional_groups: List[int] = None) -> List[Union[bool, int, float]]:
"""Builds a feature vector for an atom.
"""
features = onek_encoding_unk(atom.GetAtomicNum() - 1, ATOM_FEATURES['atomic_num']) + \
onek_encoding_unk(atom.GetTotalDegree(), ATOM_FEATURES['degree']) + \
onek_encoding_unk(atom.GetFormalCharge(), ATOM_FEATURES['formal_charge']) + \
onek_encoding_unk(int(atom.GetChiralTag()), ATOM_FEATURES['chiral_tag']) + \
onek_encoding_unk(int(atom.GetTotalNumHs()), ATOM_FEATURES['num_Hs']) + \
onek_encoding_unk(int(atom.GetHybridization()), ATOM_FEATURES['hybridization']) + \
[1 if atom.GetIsAromatic() else 0] + \
[atom.GetMass() * 0.01] # scaled to about the same range as other features
if functional_groups is not None:
features += functional_groups
return features
def initialize_weights(model: nn.Module) -> None:
"""Initializes the weights of a model in place.
"""
for param in model.parameters():
if param.dim() == 1:
nn.init.constant_(param, 0)
else:
nn.init.xavier_normal_(param)
class MPNEncoder(nn.Module):
def __init__(self, args, atom_fdim, bond_fdim):
super(MPNEncoder, self).__init__()
self.atom_fdim = atom_fdim
self.bond_fdim = bond_fdim
self.hidden_size = args.hidden_size
self.bias = args.bias
self.depth = args.depth
self.dropout = args.dropout
self.layers_per_message = 1
self.undirected = False
self.atom_messages = False
self.device = args.device
self.aggregation = args.aggregation
self.aggregation_norm = args.aggregation_norm
self.dropout_layer = nn.Dropout(p=self.dropout)
self.act_func = nn.ReLU()
self.cached_zero_vector = nn.Parameter(torch.zeros(self.hidden_size), requires_grad=False)
# Input
input_dim = self.bond_fdim
self.W_i = nn.Linear(input_dim, self.hidden_size, bias=self.bias)
w_h_input_size = self.hidden_size
# Shared weight matrix across depths (default)
self.W_h = nn.Linear(w_h_input_size, self.hidden_size, bias=self.bias)
self.W_o = nn.Linear(self.atom_fdim + self.hidden_size, self.hidden_size)
def forward(self, mol_graph):
"""Encodes a batch of molecular graphs.
"""
f_atoms, f_bonds, a2b, b2a, b2revb, a_scope, b_scope = mol_graph.get_components()
f_atoms, f_bonds, a2b, b2a, b2revb = f_atoms.to(self.device), f_bonds.to(self.device), a2b.to(self.device), b2a.to(self.device), b2revb.to(self.device)
input = self.W_i(f_bonds) # num_bonds x hidden_size
message = self.act_func(input) # num_bonds x hidden_size
# Message passing
for depth in range(self.depth - 1):
# m(a1 -> a2) = [sum_{a0 \in nei(a1)} m(a0 -> a1)] - m(a2 -> a1)
# message a_message = sum(nei_a_message) rev_message
nei_a_message = index_select_ND(message, a2b) # num_atoms x max_num_bonds x hidden
a_message = nei_a_message.sum(dim=1) # num_atoms x hidden
rev_message = message[b2revb] # num_bonds x hidden
message = a_message[b2a] - rev_message # num_bonds x hidden
message = self.W_h(message)
message = self.act_func(input + message) # num_bonds x hidden_size
message = self.dropout_layer(message) # num_bonds x hidden
a2x = a2b
nei_a_message = index_select_ND(message, a2x) # num_atoms x max_num_bonds x hidden
a_message = nei_a_message.sum(dim=1) # num_atoms x hidden
a_input = torch.cat([f_atoms, a_message], dim=1) # num_atoms x (atom_fdim + hidden)
atom_hiddens = self.act_func(self.W_o(a_input)) # num_atoms x hidden
atom_hiddens = self.dropout_layer(atom_hiddens) # num_atoms x hidden
# Readout
mol_vecs = []
for i, (a_start, a_size) in enumerate(a_scope):
if a_size == 0:
mol_vecs.append(self.cached_zero_vector)
else:
cur_hiddens = atom_hiddens.narrow(0, a_start, a_size)
mol_vec = cur_hiddens # (num_atoms, hidden_size)
if self.aggregation == 'mean':
mol_vec = mol_vec.sum(dim=0) / a_size
elif self.aggregation == 'sum':
mol_vec = mol_vec.sum(dim=0)
elif self.aggregation == 'norm':
mol_vec = mol_vec.sum(dim=0) / self.aggregation_norm
mol_vecs.append(mol_vec)
mol_vecs = torch.stack(mol_vecs, dim=0) # (num_molecules, hidden_size)
return mol_vecs # num_molecules x hidden
class MPN(nn.Module):
def __init__(self, args, atom_fdim=None, bond_fdim=None):
super(MPN, self).__init__()
self.atom_fdim = atom_fdim or get_atom_fdim()
self.bond_fdim = bond_fdim or get_bond_fdim()
self.device = args.device
self.encoder = MPNEncoder(args, self.atom_fdim, self.bond_fdim)
def forward(self, batch):
"""Encodes a batch of molecules.
"""
if type(batch[0]) != BatchMolGraph:
batch = [mol2graph(b) for b in batch]
encodings = [self.encoder(batch[0])]
output = reduce(lambda x, y: torch.cat((x, y), dim=1), encodings)
return output
class MoleculeModel(nn.Module):
def __init__(self, args, featurizer=False):
super(MoleculeModel, self).__init__()
self.classification = args.dataset_type == 'classification'
self.featurizer = featurizer
self.output_size = args.num_tasks
if self.classification:
self.sigmoid = nn.Sigmoid()
self.create_encoder(args)
self.create_ffn(args)
initialize_weights(self)
def create_encoder(self, args):
self.encoder = MPN(args)
def create_ffn(self, args):
first_linear_dim = args.hidden_size
dropout = nn.Dropout(args.dropout)
activation = nn.ReLU()
# Create FFN layers
if args.ffn_num_layers == 1:
ffn = [
dropout,
nn.Linear(first_linear_dim, self.output_size)
]
else:
ffn = [
dropout,
nn.Linear(first_linear_dim, args.ffn_hidden_size)
]
for _ in range(args.ffn_num_layers - 2):
ffn.extend([
activation,
dropout,
nn.Linear(args.ffn_hidden_size, args.ffn_hidden_size),
])
ffn.extend([
activation,
dropout,
nn.Linear(args.ffn_hidden_size, self.output_size),
])
# Create FFN model
self.ffn = nn.Sequential(*ffn)
def featurize(self, batch, features_batch=None, atom_descriptors_batch=None):
"""Computes feature vectors of the input by running the model except for the last layer.
"""
return self.ffn[:-1](self.encoder(batch, features_batch, atom_descriptors_batch))
def forward(self, batch):
output = self.ffn(self.encoder(batch))
# Don't apply sigmoid during training b/c using BCEWithLogitsLoss
if self.classification and not self.training:
output = self.sigmoid(output)
return output